10.16. Gene Explorer Tool¶
Note
EdX does not support this tool.
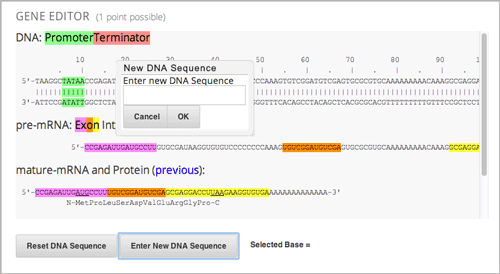
The gene explorer (GeneX), from the biology department at UMB, simulates the transcription, splicing, processing, and translation of a small hypothetical eukaryotic gene. GeneX allows learners to make specific mutations in a gene sequence, and it then calculates and displays the effects of the mutations on the mRNA and protein.
Specifically, the gene explorer does the following:
Finds the promoter and terminator.
Reads the DNA sequence to produce the pre-mRNA.
Finds the splice sites.
Splices and tails the mRNA.
Finds the start codon.
Translates the mRNA.
For more information about the gene explorer, see The Gene Explorer.
10.16.1. Gene Explorer Code¶
<problem>
<p>Make a single base pair substitution mutation in the gene below that results in a protein that is longer than the protein produced by the original gene. When you are satisfied with your change and its effect, click the <b>SUBMIT</b> button.</p>
<p>Note that a "single base pair substitution mutation" is when a single base is changed to another base; for example, changing the A at position 80 to a T. Deletions and insertions are not allowed.</p>
<script type="loncapa/python">
def genex_grader(expect,ans):
if ans=="CORRECT": return True
import json
ans=json.loads(ans)
return ans["genex_answer"]=="CORRECT"
</script>
<customresponse cfn="genex_grader">
<editageneinput width="818" height="1000" dna_sequence="TAAGGCTATAACCGAGATTGATGCCTTGTGCGATAAGGTGTGTCCCCCCCCAAAGTGTCGGATGTCGAGTGCGCGTGCAAAAAAAAACAAAGGCGAGGACCTTAAGAAGGTGTGAGGGGGCGCTCGAT" genex_dna_sequence="TAAGGCTATAACCGAGATTGATGCCTTGTGCGATAAGGTGTGTCCCCCCCCAAAGTGTCGGATGTCGAGTGCGCGTGCAAAAAAAAACAAAGGCGAGGACCTTAAGAAGGTGTGAGGGGGCGCTCGAT" genex_problem_number="2"/>
</customresponse>
</problem>
In this code:
width and height specify the dimensions of the application, in pixels.
genex_dna_sequence is the default DNA sequence that appears when the problem opens.
dna_sequence contains the application’s state and the learner’s answer. This value must be the same as genex_dna_sequence.
genex_problem_number specifies the number of the problem. This number is based on the five gene editor problems in the MITx 7.00x course–for example, if you want this problem to look like the second gene editor problem in the 7.00x course, you would set the genex_problem_number value to 2. The number must be 1, 2, 3, 4, or 5.